Welcome to Gan Jing World!
seo.Headlines
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Trending Videos
The UFO Crash at Roswell

Category:cat33
Channel:Filmhub
Duration:3287seconds
Hashtags:
Like Counts:15
View Counts:106
Status:
Type:Movie
Bicycle Handsome Factory Deck Review

Category:cat8
Channel:Shuffle Up and Deal
Duration:161seconds
Hashtags:
Like Counts:1
View Counts:18
Status:
Type:Video
Animals Reunited With Owners After Years! D Animals Reunited
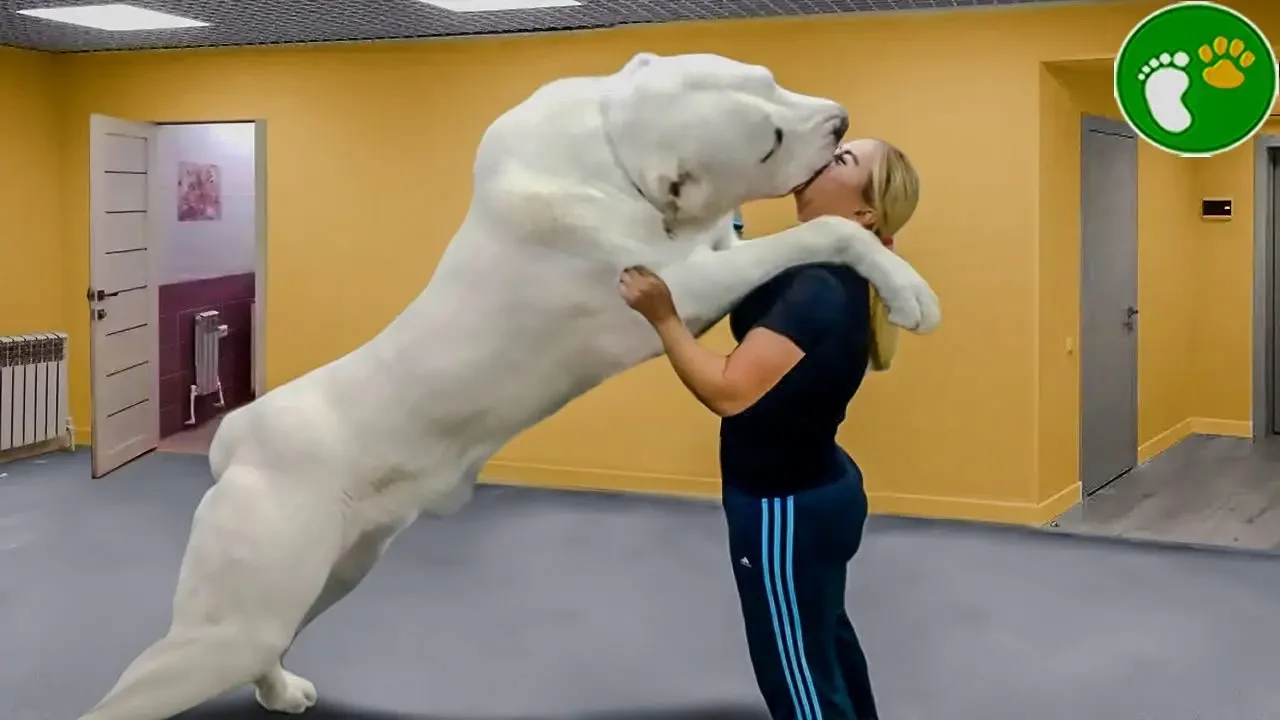

Category:cat8
Channel:D - Animals Reunited
Duration:4663seconds
Hashtags:#29,#animalreunions,#animalsreunitedwithowners
Like Counts:11
View Counts:164
Status:
Type:Video
HangTown: Cartoonist Daxiong's Latest Music Video | Exclusive to Gan Jing World | Must-watch

Category:cat17
Channel:Da Xiong Originals
Duration:643seconds
Hashtags:#2024必聽,#乾净世界獨家,#大雄,#大雄原創,#天朝上國,#官方,#流行音樂,#熱播,#熱門歌曲,#獨家
Like Counts:5828
View Counts:131593
Status:
Type:Video
Quick yogurt cake is one of the most successful cakes you can make, which melts in your mouth

Category:cat9
Channel:Home Made Yum Yum
Duration:240seconds
Hashtags:#Lemoncake,#blueberry,#blueberryyogurtcake,#cupcakeswithmary,#greekyogurtcakerecipe,#lemonblueberrycake,#lemoncake,#lemonyogurtcakeinagarten,#makevanillalemoncakeeverydayinafewminutes,#muffinsrecipe,#vanillacake,#vanillalemoncake,#yogurtbread,#yogurtcake3ingredients,#yogurtcakerecipeeasy
Like Counts:1
View Counts:8
Status:
Type:Video
The Alexander Show

Category:cat33
Channel:Filmhub
Duration:0seconds
Hashtags:
Like Counts:4
View Counts:58
Status:
Type:TVSeries
The Most Modern Agriculture Machines That Are At Another Level , How To Harvest Oranges In Farm ▶3
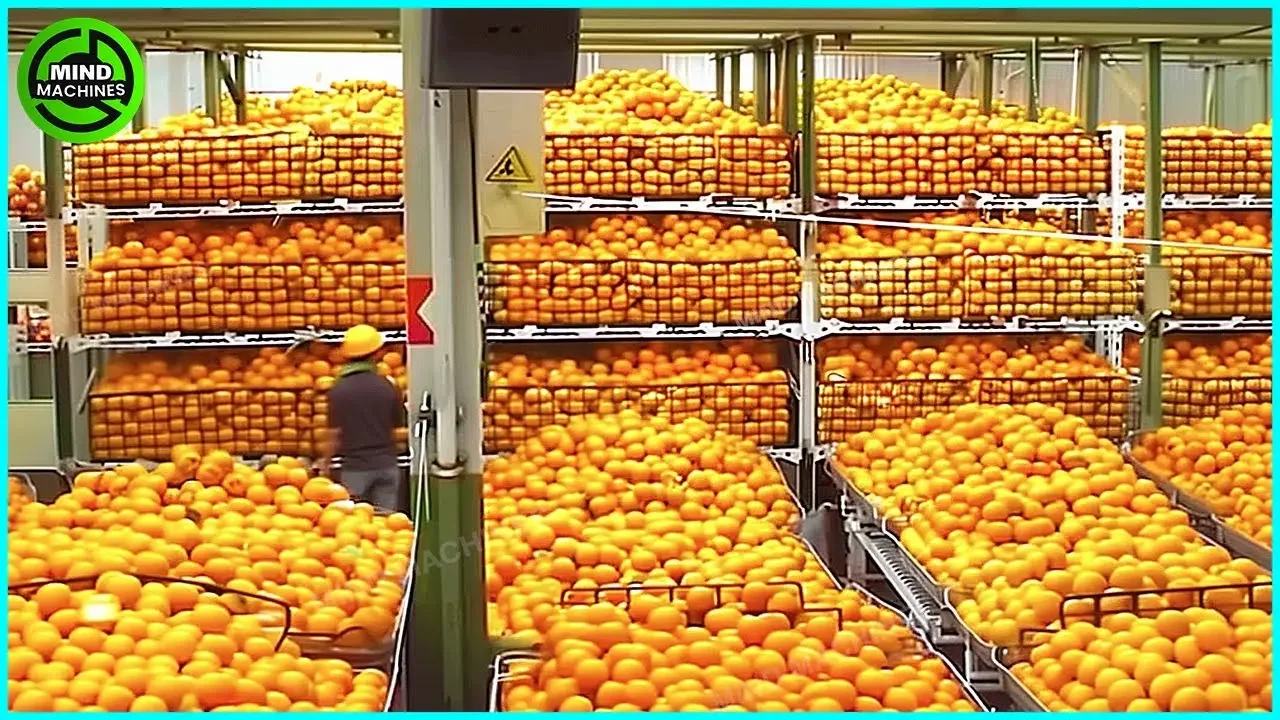

Category:cat23
Channel:MIND MACHINES
Duration:2110seconds
Hashtags:#agriculturemachines,#howtoharvest,#machines,#modernagriculturemachines
Like Counts:2
View Counts:12
Status:
Type:Video
How to Get the BEST Redemptions with Capital One Miles! (and AVOID the Worst)

Category:cat15
Channel:Venture with Jazzy
Duration:743seconds
Hashtags:
Like Counts:2
View Counts:15
Status:
Type:Video
New York Times Development of Hit Piece on Shen Yun Akin to Chinese Regime’s Propaganda: Reporter

Category:cat19
Channel:NTD
Duration:279seconds
Hashtags:
Like Counts:116
View Counts:959
Status:
Type:Video
Discovering the Traditional Bel Canto, Episode 6

Category:cat2
Channel:Shen Yun Creations
Duration:539seconds
Hashtags:
Like Counts:23
View Counts:1423
Status:
Type:Video
Traditional Chinese Tales:Integrity Brings Heavenly Assistance

Category:cat7
Channel:Jing Di Academy - A GanJingWorld.com Exclusive
Duration:270seconds
Hashtags:
Like Counts:22
View Counts:178
Status:
Type:Video
5,000 Years of Heroes - Han Xin

Category:cat33
Channel:True Legend Studio
Duration:0seconds
Hashtags:
Like Counts:117
View Counts:485
Status:
Type:TVSeries
seo.trendingStartKey
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Category:
Channel:
Duration:undefinedseconds
Hashtags:
Like Counts:
View Counts:
Status:
Type:
Top Stories
House Advance Foreign Aid and TikTok Bills

Category:cat19
Channel:The Beltway Pundit
Duration:32seconds
Hashtags:
Like Counts:4
View Counts:91
Status:
Type:Video
NTD Evening News Full Broadcast (April 19)

Category:cat19
Channel:NTD
Duration:0seconds
Hashtags:
Like Counts:5
View Counts:9
Status:LIVE
Type:Live
Man Sets Himself on Fire Outside Trump Trial Courthouse

Category:cat19
Channel:The Latest
Duration:201seconds
Hashtags:
Like Counts:2
View Counts:17
Status:
Type:Video
Rep. Jeffries Blasts GOP After House Advanced Ukraine Aid

Category:cat19
Channel:Pure News
Duration:74seconds
Hashtags:
Like Counts:0
View Counts:16
Status:
Type:Video
FBI Director Christopher Wray Remarks on China's Threat

Category:cat19
Channel:The Rundown
Duration:1389seconds
Hashtags:
Like Counts:1
View Counts:3
Status:
Type:Video
China Orders Apple to Remove Popular Apps

Category:cat19
Channel:Main Street
Duration:56seconds
Hashtags:
Like Counts:0
View Counts:9
Status:
Type:Video
A Divided House GOP Jeopardizes Johnson's Speakership

Category:cat19
Channel:News Digest
Duration:106seconds
Hashtags:
Like Counts:24
View Counts:94
Status:
Type:Video
US Vetoes Palestine Membership into the United Nations

Category:cat19
Channel:Live News Network
Duration:59seconds
Hashtags:
Like Counts:1
View Counts:7
Status:
Type:Video
US Officials: Israel hits Iran with limited military strike
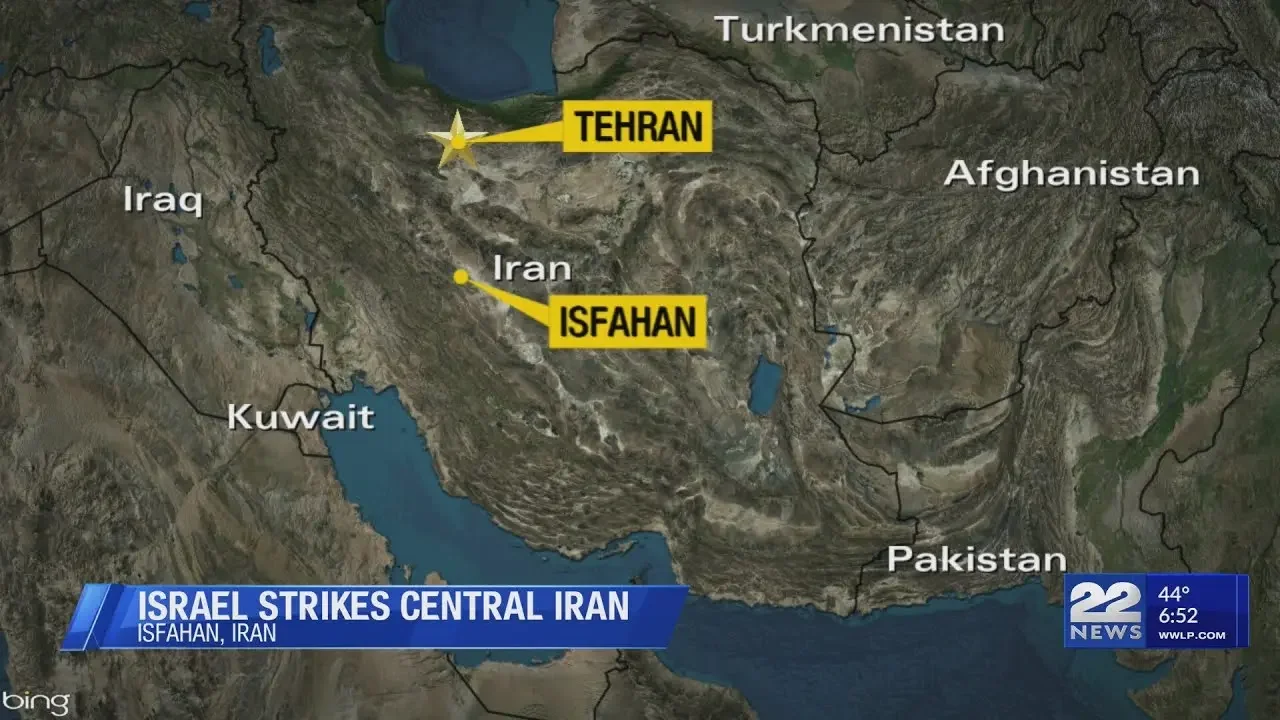
Category:cat19
Channel:Western Massachusetts News, Weather & Sports
Duration:108seconds
Hashtags:
Like Counts:4
View Counts:12
Status:
Type:Video
Close call collision narrowly avoided after 2 jets were cleared onto the same runway

Category:cat19
Channel:6abc Philadelphia
Duration:61seconds
Hashtags:
Like Counts:0
View Counts:17
Status:
Type:Video
New Taylor Swift album to release Friday
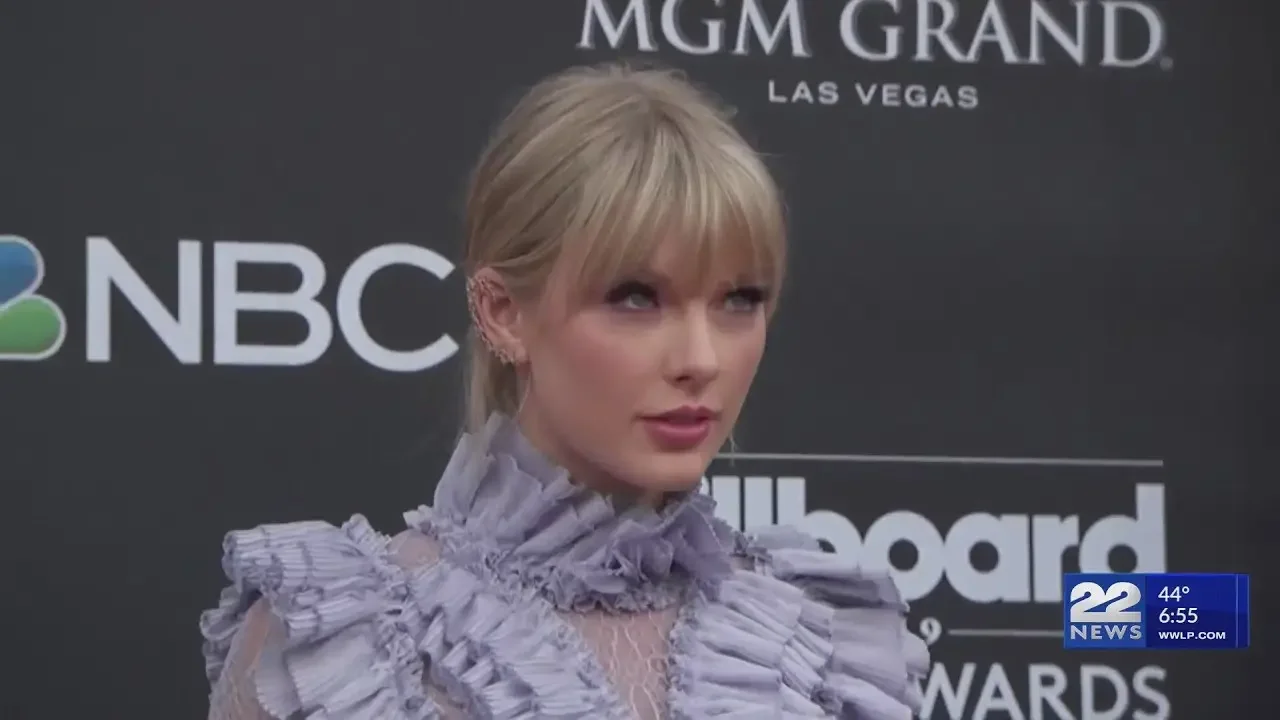

Category:cat19
Channel:Western Massachusetts News, Weather & Sports
Duration:39seconds
Hashtags:
Like Counts:0
View Counts:9
Status:
Type:Video
US Air Force Grilled Over F-35 Program

Category:cat19
Channel:Pure News
Duration:311seconds
Hashtags:
Like Counts:14
View Counts:54
Status:
Type:Video
Kenya's Military Chief, 9 Others Killed in Helicopter Crash

Category:cat19
Channel:The Latest
Duration:478seconds
Hashtags:
Like Counts:18
View Counts:59
Status:
Type:Video